an active gene leads to the production of what? (one word)
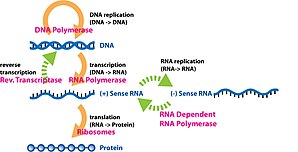
Cistron expression is the procedure by which information from a gene is used in the synthesis of a functional cistron production that enables information technology to produce stop products, protein or non-coding RNA, and ultimately affect a phenotype, every bit the last effect. These products are oftentimes proteins, only in non-protein-coding genes such as transfer RNA (tRNA) and pocket-sized nuclear RNA (snRNA), the product is a functional non-coding RNA. Gene expression is summarized in the key dogma of molecular biology first formulated by Francis Crick in 1958,[1] further developed in his 1970 article,[2] and expanded by the subsequent discoveries of opposite transcription[3] [4] [5] and RNA replication.[half-dozen]
The process of gene expression is used by all known life—eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea), and utilized by viruses—to generate the macromolecular mechanism for life.
In genetics, gene expression is the most fundamental level at which the genotype gives ascent to the phenotype, i.e. observable trait. The genetic data stored in Dna represents the genotype, whereas the phenotype results from the "estimation" of that data. Such phenotypes are often expressed by the synthesis of proteins that command the organism's construction and development, or that act equally enzymes catalyzing specific metabolic pathways.
All steps in the factor expression procedure may be modulated (regulated), including the transcription, RNA splicing, translation, and mail service-translational modification of a poly peptide. Regulation of gene expression gives command over the timing, location, and corporeality of a given gene production (protein or ncRNA) present in a prison cell and tin can have a profound effect on the cellular construction and function. Regulation of gene expression is the basis for cellular differentiation, evolution, morphogenesis and the versatility and adaptability of whatever organism. Cistron regulation may therefore serve equally a substrate for evolutionary modify.
Machinery [edit]
Transcription [edit]

The process of transcription is carried out by RNA polymerase (RNAP), which uses Deoxyribonucleic acid (black) equally a template and produces RNA (bluish).
The production of a RNA copy from a DNA strand is called transcription, and is performed past RNA polymerases, which add together one ribonucleotide at a time to a growing RNA strand as per the complementarity constabulary of the nucleotide bases. This RNA is complementary to the template 3′ → five′ Deoxyribonucleic acid strand,[7] with the exception that thymines (T) are replaced with uracils (U) in the RNA.
In prokaryotes, transcription is carried out by a single type of RNA polymerase, which needs to bind a Dna sequence called a Pribnow box with the help of the sigma gene protein (σ cistron) to start transcription. In eukaryotes, transcription is performed in the nucleus by 3 types of RNA polymerases, each of which needs a special Dna sequence chosen the promoter and a set of DNA-binding proteins—transcription factors—to initiate the procedure (see regulation of transcription beneath). RNA polymerase I is responsible for transcription of ribosomal RNA (rRNA) genes. RNA polymerase Two (Politico II) transcribes all poly peptide-coding genes simply also some non-coding RNAs (e.g., snRNAs, snoRNAs or long non-coding RNAs). RNA polymerase III transcribes 5S rRNA, transfer RNA (tRNA) genes, and some small non-coding RNAs (e.g., 7SK). Transcription ends when the polymerase encounters a sequence called the terminator.
mRNA processing [edit]
While transcription of prokaryotic protein-coding genes creates messenger RNA (mRNA) that is ready for translation into protein, transcription of eukaryotic genes leaves a master transcript of RNA (pre-RNA), which outset has to undergo a serial of modifications to become a mature RNA. Types and steps involved in the maturation processes vary betwixt coding and non-coding preRNAs; i.east. even though preRNA molecules for both mRNA and tRNA undergo splicing, the steps and machinery involved are different.[8] The processing of non-coding RNA is described below (non-coring RNA maturation).
The processing of premRNA include 5′ capping, which is fix of enzymatic reactions that add 7-methylguanosine (msevenG) to the 5′ end of pre-mRNA and thus protect the RNA from deposition by exonucleases. The msevenGrand cap is so bound by cap binding complex heterodimer (CBC20/CBC80), which aids in mRNA export to cytoplasm and also protect the RNA from decapping.
Some other modification is 3′ cleavage and polyadenylation. They occur if polyadenylation indicate sequence (five′- AAUAAA-3′) is present in pre-mRNA, which is usually between protein-coding sequence and terminator. The pre-mRNA is kickoff cleaved and then a series of ~200 adenines (A) are added to form poly(A) tail, which protects the RNA from deposition. The poly(A) tail is bound by multiple poly(A)-binding proteins (PABPs) necessary for mRNA export and translation re-initiation. In the inverse process of deadenylation, poly(A) tails are shortened by the CCR4-Not 3′-5′ exonuclease, which ofttimes leads to full transcript disuse.
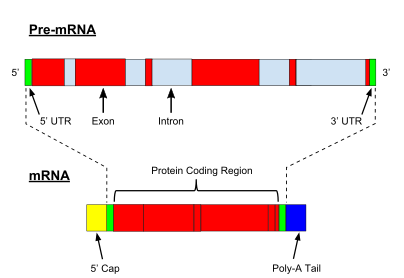
Analogy of exons and introns in pre-mRNA and the germination of mature mRNA by splicing. The UTRs (in greenish) are non-coding parts of exons at the ends of the mRNA.
A very important modification of eukaryotic pre-mRNA is RNA splicing. The majority of eukaryotic pre-mRNAs consist of alternating segments called exons and introns. During the procedure of splicing, an RNA-protein catalytical complex known as spliceosome catalyzes two transesterification reactions, which remove an intron and release it in form of lariat construction, and and then splice neighbouring exons together. In certain cases, some introns or exons can exist either removed or retained in mature mRNA. This so-chosen alternative splicing creates serial of different transcripts originating from a single cistron. Because these transcripts tin can be potentially translated into dissimilar proteins, splicing extends the complexity of eukaryotic gene expression and the size of a species proteome.
All-encompassing RNA processing may be an evolutionary advantage made possible by the nucleus of eukaryotes. In prokaryotes, transcription and translation happen together, whilst in eukaryotes, the nuclear membrane separates the two processes, giving time for RNA processing to occur.
Non-coding RNA maturation [edit]
In well-nigh organisms not-coding genes (ncRNA) are transcribed equally precursors that undergo further processing. In the case of ribosomal RNAs (rRNA), they are ofttimes transcribed as a pre-rRNA that contains i or more rRNAs. The pre-rRNA is cleaved and modified (2′-O-methylation and pseudouridine formation) at specific sites by approximately 150 unlike pocket-size nucleolus-restricted RNA species, chosen snoRNAs. SnoRNAs associate with proteins, forming snoRNPs. While snoRNA part basepair with the target RNA and thus position the modification at a precise site, the poly peptide part performs the catalytical reaction. In eukaryotes, in particular a snoRNP called RNase, MRP cleaves the 45S pre-rRNA into the 28S, five.8S, and 18S rRNAs. The rRNA and RNA processing factors grade large aggregates called the nucleolus.[9]
In the case of transfer RNA (tRNA), for example, the 5′ sequence is removed by RNase P,[10] whereas the 3′ end is removed by the tRNase Z enzyme[xi] and the non-templated 3′ CCA tail is added by a nucleotidyl transferase.[12] In the case of micro RNA (miRNA), miRNAs are first transcribed equally main transcripts or pri-miRNA with a cap and poly-A tail and candy to brusque, 70-nucleotide stem-loop structures known as pre-miRNA in the cell nucleus past the enzymes Drosha and Pasha. Later being exported, it is then processed to mature miRNAs in the cytoplasm by interaction with the endonuclease Dicer, which also initiates the formation of the RNA-induced silencing complex (RISC), composed of the Argonaute protein.
Even snRNAs and snoRNAs themselves undergo series of modification before they become part of functional RNP circuitous. This is done either in the nucleoplasm or in the specialized compartments called Cajal bodies. Their bases are methylated or pseudouridinilated by a group of minor Cajal body-specific RNAs (scaRNAs), which are structurally similar to snoRNAs.
RNA export [edit]
In eukaryotes most mature RNA must be exported to the cytoplasm from the nucleus. While some RNAs function in the nucleus, many RNAs are transported through the nuclear pores and into the cytosol.[xiii] Export of RNAs requires clan with specific proteins known as exportins. Specific exportin molecules are responsible for the export of a given RNA type. mRNA transport also requires the correct association with Exon Junction Complex (EJC), which ensures that correct processing of the mRNA is completed before consign. In some cases RNAs are additionally transported to a specific function of the cytoplasm, such as a synapse; they are then towed by motor proteins that bind through linker proteins to specific sequences (called "zipcodes") on the RNA.[14]
Translation [edit]
For some RNA (not-coding RNA) the mature RNA is the final gene product.[15] In the case of messenger RNA (mRNA) the RNA is an data carrier coding for the synthesis of i or more proteins. mRNA conveying a single poly peptide sequence (common in eukaryotes) is monocistronic whilst mRNA carrying multiple protein sequences (mutual in prokaryotes) is known as polycistronic.
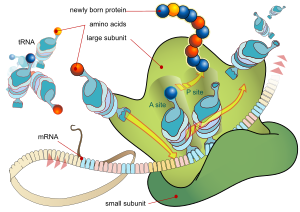
During the translation, tRNA charged with amino acid enters the ribosome and aligns with the correct mRNA triplet. Ribosome then adds amino acid to growing poly peptide concatenation.
Every mRNA consists of three parts: a 5′ untranslated region (5′UTR), a poly peptide-coding region or open reading frame (ORF), and a 3′ untranslated region (3′UTR). The coding region carries information for protein synthesis encoded past the genetic code to form triplets. Each triplet of nucleotides of the coding region is called a codon and corresponds to a bounden site complementary to an anticodon triplet in transfer RNA. Transfer RNAs with the same anticodon sequence always carry an identical type of amino acid. Amino acids are then chained together by the ribosome co-ordinate to the order of triplets in the coding region. The ribosome helps transfer RNA to bind to messenger RNA and takes the amino acid from each transfer RNA and makes a construction-less protein out of it.[sixteen] [17] Each mRNA molecule is translated into many protein molecules, on average ~2800 in mammals.[xviii] [19]
In prokaryotes translation by and large occurs at the point of transcription (co-transcriptionally), frequently using a messenger RNA that is still in the process of beingness created. In eukaryotes translation tin occur in a variety of regions of the cell depending on where the protein existence written is supposed to be. Major locations are the cytoplasm for soluble cytoplasmic proteins and the membrane of the endoplasmic reticulum for proteins that are for export from the cell or insertion into a prison cell membrane. Proteins that are supposed to be expressed at the endoplasmic reticulum are recognised function-way through the translation process. This is governed by the point recognition particle—a protein that binds to the ribosome and directs it to the endoplasmic reticulum when it finds a signal peptide on the growing (nascent) amino acid chain.[20]
Folding [edit]
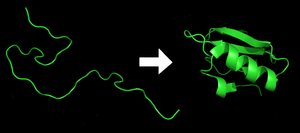
Protein earlier (left) and afterward (right) folding
Each poly peptide exists as an unfolded polypeptide or random coil when translated from a sequence of mRNA into a linear chain of amino acids. This polypeptide lacks any developed three-dimensional construction (the left mitt side of the neighboring figure). The polypeptide then folds into its characteristic and functional three-dimensional structure from a random coil.[21] Amino acids collaborate with each other to produce a well-divers 3-dimensional structure, the folded protein (the right mitt side of the figure) known as the native state. The resulting three-dimensional construction is adamant by the amino acid sequence (Anfinsen'southward dogma).[22]
The correct iii-dimensional structure is essential to part, although some parts of functional proteins may remain unfolded.[23] Failure to fold into the intended shape usually produces inactive proteins with different backdrop including toxic prions. Several neurodegenerative and other diseases are believed to result from the accumulation of misfolded proteins.[24] Many allergies are caused by the folding of the proteins, for the allowed system does not produce antibodies for certain protein structures.[25]
Enzymes called chaperones assist the newly formed protein to attain (fold into) the 3-dimensional structure it needs to function.[26] Similarly, RNA chaperones help RNAs attain their functional shapes.[27] Assisting protein folding is one of the main roles of the endoplasmic reticulum in eukaryotes.
Translocation [edit]
Secretory proteins of eukaryotes or prokaryotes must exist translocated to enter the secretory pathway. Newly synthesized proteins are directed to the eukaryotic Sec61 or prokaryotic SecYEG translocation channel by signal peptides. The efficiency of protein secretion in eukaryotes is very dependent on the signal peptide which has been used.[28]
Protein ship [edit]
Many proteins are destined for other parts of the cell than the cytosol and a wide range of signalling sequences or (signal peptides) are used to direct proteins to where they are supposed to be. In prokaryotes this is normally a simple process due to express compartmentalisation of the cell. However, in eukaryotes there is a great variety of different targeting processes to ensure the protein arrives at the correct organelle.
Not all proteins remain within the cell and many are exported, for example, digestive enzymes, hormones and extracellular matrix proteins. In eukaryotes the export pathway is well developed and the primary mechanism for the export of these proteins is translocation to the endoplasmic reticulum, followed by transport via the Golgi apparatus.[29] [thirty]
Regulation of gene expression [edit]

Regulation of gene expression is the command of the amount and timing of appearance of the functional product of a gene. Command of expression is vital to allow a cell to produce the gene products information technology needs when it needs them; in turn, this gives cells the flexibility to adapt to a variable environment, external signals, damage to the cell, and other stimuli. More mostly, gene regulation gives the prison cell control over all structure and function, and is the basis for cellular differentiation, morphogenesis and the versatility and adaptability of whatsoever organism.
Numerous terms are used to describe types of genes depending on how they are regulated; these include:
- A constitutive cistron is a cistron that is transcribed continually as opposed to a facultative cistron, which is simply transcribed when needed.
- A housekeeping gene is a gene that is required to maintain basic cellular role and and so is typically expressed in all cell types of an organism. Examples include actin, GAPDH and ubiquitin. Some housekeeping genes are transcribed at a relatively constant rate and these genes can be used as a reference signal in experiments to mensurate the expression rates of other genes.
- A facultative gene is a gene just transcribed when needed every bit opposed to a constitutive gene.
- An inducible factor is a factor whose expression is either responsive to environmental change or dependent on the position in the cell wheel.
Any step of gene expression may exist modulated, from the Dna-RNA transcription step to post-translational modification of a protein. The stability of the final gene product, whether it is RNA or protein, also contributes to the expression level of the cistron—an unstable production results in a low expression level. In general gene expression is regulated through changes[31] in the number and type of interactions between molecules[32] that collectively influence transcription of DNA[33] and translation of RNA.[34]
Some uncomplicated examples of where gene expression is important are:
- Control of insulin expression then it gives a indicate for blood glucose regulation.
- X chromosome inactivation in female person mammals to preclude an "overdose" of the genes it contains.
- Cyclin expression levels control progression through the eukaryotic cell cycle.
Transcriptional regulation [edit]

When lactose is present in a prokaryote, it acts equally an inducer and inactivates the repressor so that the genes for lactose metabolism can be transcribed.
Regulation of transcription tin be cleaved down into iii main routes of influence; genetic (direct interaction of a control cistron with the factor), modulation interaction of a control factor with the transcription mechanism and epigenetic (not-sequence changes in DNA structure that influence transcription).

Direct interaction with Deoxyribonucleic acid is the simplest and the most direct method past which a protein changes transcription levels. Genes often have several protein binding sites around the coding region with the specific function of regulating transcription. There are many classes of regulatory DNA binding sites known every bit enhancers, insulators and silencers. The mechanisms for regulating transcription are varied, from blocking key binding sites on the Deoxyribonucleic acid for RNA polymerase to acting as an activator and promoting transcription past assisting RNA polymerase binding.
The activeness of transcription factors is further modulated by intracellular signals causing protein post-translational modification including phosphorylated, acetylated, or glycosylated. These changes influence a transcription factor'due south ability to demark, directly or indirectly, to promoter DNA, to recruit RNA polymerase, or to favor elongation of a newly synthesized RNA molecule.
The nuclear membrane in eukaryotes allows further regulation of transcription factors by the elapsing of their presence in the nucleus, which is regulated past reversible changes in their structure and by bounden of other proteins.[35] Environmental stimuli or endocrine signals[36] may cause modification of regulatory proteins[37] eliciting cascades of intracellular signals,[38] which result in regulation of gene expression.
More recently it has become credible that there is a significant influence of non-Deoxyribonucleic acid-sequence specific effects on transcription. These effects are referred to every bit epigenetic and involve the higher club structure of DNA, non-sequence specific DNA binding proteins and chemic modification of DNA. In general epigenetic effects change the accessibility of DNA to proteins and then modulate transcription.

In eukaryotes, DNA is organized in grade of nucleosomes. Note how the Deoxyribonucleic acid (blue and green) is tightly wrapped around the protein core made of histone octamer (ribbon coils), restricting access to the Dna. From PDB: 1KX5.
In eukaryotes the structure of chromatin, controlled past the histone lawmaking, regulates access to DNA with meaning impacts on the expression of genes in euchromatin and heterochromatin areas.
Enhancers, transcription factors, mediator circuitous and Deoxyribonucleic acid loops in mammalian transcription [edit]
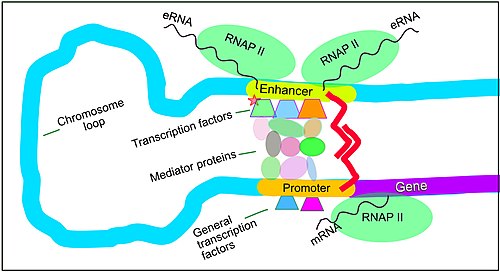
Regulation of transcription in mammals. An active enhancer regulatory region is enabled to interact with the promoter region of its target factor by formation of a chromosome loop. This can initiate messenger RNA (mRNA) synthesis by RNA polymerase II (RNAP Two) bound to the promoter at the transcription starting time site of the gene. The loop is stabilized by one architectural poly peptide anchored to the enhancer and one anchored to the promoter and these proteins are joined to grade a dimer (red zigzags). Specific regulatory transcription factors bind to Dna sequence motifs on the enhancer. Full general transcription factors bind to the promoter. When a transcription factor is activated by a betoken (here indicated as phosphorylation shown by a pocket-size red star on a transcription factor on the enhancer) the enhancer is activated and can at present activate its target promoter. The active enhancer is transcribed on each strand of Dna in contrary directions by bound RNAP IIs. Mediator (a complex consisting of about 26 proteins in an interacting structure) communicates regulatory signals from the enhancer Deoxyribonucleic acid-bound transcription factors to the promoter.
Gene expression in mammals is regulated by many cis-regulatory elements, including cadre promoters and promoter-proximal elements that are located almost the transcription start sites of genes, upstream on the DNA (towards the 5' region of the sense strand). Other of import cis-regulatory modules are localized in DNA regions that are distant from the transcription start sites. These include enhancers, silencers, insulators and tethering elements.[39] Enhancers and their associated transcription factors have a leading role in the regulation of factor expression.[40]
Enhancers are genome regions that regulate genes. Enhancers control cell-type-specific gene expression programs, most often by looping through long distances to come in concrete proximity with the promoters of their target genes.[41] Multiple enhancers, each often tens or hundred of thousands of nucleotides afar from their target genes, loop to their target gene promoters and coordinate with each other to control gene expression.[41]
The analogy shows an enhancer looping around to come into proximity with the promoter of a target cistron. The loop is stabilized by a dimer of a connector poly peptide (e.yard. dimer of CTCF or YY1). Ane member of the dimer is anchored to its binding motif on the enhancer and the other member is anchored to its binding motif on the promoter (represented by the scarlet zigzags in the analogy).[42] Several jail cell function-specific transcription factors (among the about i,600 transcription factors in a homo prison cell)[43] generally bind to specific motifs on an enhancer.[44] A minor combination of these enhancer-spring transcription factors, when brought close to a promoter by a Dna loop, govern transcription level of the target gene. Mediator (a complex usually consisting of most 26 proteins in an interacting structure) communicates regulatory signals from enhancer DNA-bound transcription factors directly to the RNA polymerase II (pol 2) enzyme jump to the promoter.[45]
Enhancers, when agile, are generally transcribed from both strands of DNA with RNA polymerases interim in ii different directions, producing ii eRNAs as illustrated in the figure.[46] An inactive enhancer may exist spring by an inactive transcription factor. Phosphorylation of the transcription gene may actuate information technology and that activated transcription factor may then activate the enhancer to which it is bound (see small red star representing phosphorylation of transcription gene bound to enhancer in the illustration).[47] An activated enhancer begins transcription of its RNA before activating transcription of messenger RNA from its target gene.[48]
DNA methylation and demethylation in transcriptional regulation [edit]

Deoxyribonucleic acid methylation is the addition of a methyl group to the DNA that happens at cytosine. The epitome shows a cytosine single ring base and a methyl grouping added on to the 5 carbon. In mammals, Dna methylation occurs almost exclusively at a cytosine that is followed by a guanine.
DNA methylation is a widespread mechanism for epigenetic influence on gene expression and is seen in bacteria and eukaryotes and has roles in heritable transcription silencing and transcription regulation. Methylation nigh often occurs on a cytosine (see Figure). Methylation of cytosine primarily occurs in dinucleotide sequences where a cytosine is followed by a guanine, a CpG site. The number of CpG sites in the human being genome is nigh 28 million.[49] Depending on the type of prison cell, about lxx% of the CpG sites accept a methylated cytosine.[50]
Methylation of cytosine in DNA has a major role in regulating cistron expression. Methylation of CpGs in a promoter region of a gene usually represses cistron transcription[51] while methylation of CpGs in the body of a factor increases expression.[52] TET enzymes play a central function in demethylation of methylated cytosines. Demethylation of CpGs in a gene promoter by TET enzyme activity increases transcription of the cistron.[53]
Transcriptional regulation in learning and memory [edit]
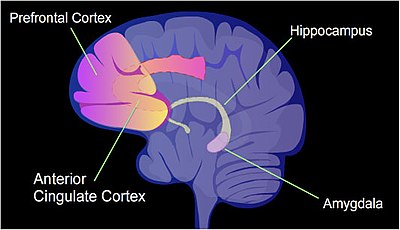
The identified areas of the homo encephalon are involved in memory germination.
In a rat, contextual fear workout (Chlorofluorocarbon) is a painful learning experience. Just ane episode of Chlorofluorocarbon tin result in a life-long fearful retention.[54] After an episode of Cfc, cytosine methylation is altered in the promoter regions of nearly nine.17% of all genes in the hippocampus neuron DNA of a rat.[55] The hippocampus is where new memories are initially stored. After Cfc most 500 genes have increased transcription (often due to demethylation of CpG sites in a promoter region) and about 1,000 genes have decreased transcription (often due to newly formed five-methylcytosine at CpG sites in a promoter region). The pattern of induced and repressed genes within neurons appears to provide a molecular ground for forming the beginning transient memory of this training event in the hippocampus of the rat brain.[55]
In detail, the brain-derived neurotrophic gene factor (BDNF) is known as a "learning gene."[56] Later CFC there was upregulation of BDNF factor expression, related to decreased CpG methylation of certain internal promoters of the cistron, and this was correlated with learning.[56]
Transcriptional regulation in cancer [edit]
The majority of gene promoters contain a CpG island with numerous CpG sites.[57] When many of a gene's promoter CpG sites are methylated the cistron becomes silenced.[58] Colorectal cancers typically accept iii to 6 commuter mutations and 33 to 66 hitchhiker or passenger mutations.[59] However, transcriptional silencing may be of more than importance than mutation in causing progression to cancer. For example, in colorectal cancers near 600 to 800 genes are transcriptionally silenced past CpG island methylation (see regulation of transcription in cancer). Transcriptional repression in cancer can also occur by other epigenetic mechanisms, such as altered expression of microRNAs.[60] In breast cancer, transcriptional repression of BRCA1 may occur more ofttimes by over-expressed microRNA-182 than by hypermethylation of the BRCA1 promoter (see Low expression of BRCA1 in chest and ovarian cancers).
Post-transcriptional regulation [edit]
In eukaryotes, where consign of RNA is required earlier translation is possible, nuclear export is idea to provide additional control over factor expression. All ship in and out of the nucleus is via the nuclear pore and ship is controlled by a wide range of importin and exportin proteins.
Expression of a gene coding for a protein is only possible if the messenger RNA carrying the code survives long plenty to be translated. In a typical cell, an RNA molecule is only stable if specifically protected from degradation. RNA degradation has particular importance in regulation of expression in eukaryotic cells where mRNA has to travel significant distances earlier being translated. In eukaryotes, RNA is stabilised past sure post-transcriptional modifications, specially the v′ cap and poly-adenylated tail.
Intentional degradation of mRNA is used not just as a defence mechanism from foreign RNA (unremarkably from viruses) simply also as a route of mRNA destabilisation. If an mRNA molecule has a complementary sequence to a pocket-sized interfering RNA and then it is targeted for destruction via the RNA interference pathway.
3 prime untranslated regions and microRNAs [edit]
3 prime untranslated regions (three′UTRs) of messenger RNAs (mRNAs) often contain regulatory sequences that post-transcriptionally influence gene expression. Such 3′-UTRs often comprise both bounden sites for microRNAs (miRNAs) as well as for regulatory proteins. By binding to specific sites within the 3′-UTR, miRNAs tin can decrease cistron expression of various mRNAs by either inhibiting translation or straight causing degradation of the transcript. The 3′-UTR too may have silencer regions that bind repressor proteins that inhibit the expression of a mRNA.
The 3′-UTR often contains microRNA response elements (MREs). MREs are sequences to which miRNAs bind. These are prevalent motifs within three′-UTRs. Among all regulatory motifs within the 3′-UTRs (due east.g. including silencer regions), MREs make up about half of the motifs.
As of 2014, the miRBase web site,[61] an archive of miRNA sequences and annotations, listed 28,645 entries in 233 biologic species. Of these, 1,881 miRNAs were in annotated homo miRNA loci. miRNAs were predicted to take an boilerplate of most four hundred target mRNAs (affecting expression of several hundred genes).[62] Friedman et al.[62] estimate that >45,000 miRNA target sites within human mRNA 3′UTRs are conserved above groundwork levels, and >60% of man protein-coding genes have been under selective force per unit area to maintain pairing to miRNAs.
Direct experiments testify that a single miRNA can reduce the stability of hundreds of unique mRNAs.[63] Other experiments bear witness that a single miRNA may repress the production of hundreds of proteins, simply that this repression oft is relatively balmy (less than 2-fold).[64] [65]
The effects of miRNA dysregulation of factor expression seem to be important in cancer.[66] For instance, in gastrointestinal cancers, nine miRNAs have been identified equally epigenetically altered and effective in down regulating DNA repair enzymes.[67]
The effects of miRNA dysregulation of cistron expression also seem to exist important in neuropsychiatric disorders, such as schizophrenia, bipolar disorder, major depression, Parkinson's illness, Alzheimer's disease and autism spectrum disorders.[68] [69]
Translational regulation [edit]
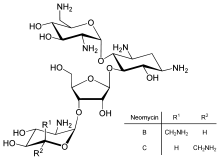
Neomycin is an instance of a small-scale molecule that reduces expression of all poly peptide genes inevitably leading to cell death; it thus acts every bit an antibiotic.
Direct regulation of translation is less prevalent than command of transcription or mRNA stability but is occasionally used. Inhibition of poly peptide translation is a major target for toxins and antibiotics, so they tin can kill a cell by overriding its normal gene expression command. Protein synthesis inhibitors include the antibody neomycin and the toxin ricin.
Post-translational modifications [edit]
Post-translational modifications (PTMs) are covalent modifications to proteins. Like RNA splicing, they help to significantly diversify the proteome. These modifications are unremarkably catalyzed by enzymes. Additionally, processes like covalent additions to amino acid side concatenation residues can often be reversed by other enzymes. However, some, like the proteolytic cleavage of the protein courage, are irreversible.[70]
PTMs play many important roles in the cell.[71] For example, phosphorylation is primarily involved in activating and deactivating proteins and in signaling pathways.[72] PTMs are involved in transcriptional regulation: an important function of acetylation and methylation is histone tail modification, which alters how accessible Deoxyribonucleic acid is for transcription.[70] They tin besides be seen in the immune system, where glycosylation plays a key role.[73] One blazon of PTM tin can initiate some other type of PTM, every bit can be seen in how ubiquitination tags proteins for degradation through proteolysis.[70] Proteolysis, other than beingness involved in breaking downwardly proteins, is as well of import in activating and deactivating them, and in regulating biological processes such every bit DNA transcription and cell decease.[74]
Measurement [edit]
Measuring factor expression is an important part of many life sciences, as the power to quantify the level at which a particular gene is expressed within a cell, tissue or organism can provide a lot of valuable information. For instance, measuring cistron expression can:
- Identify viral infection of a cell (viral poly peptide expression).
- Determine an individual's susceptibility to cancer (oncogene expression).
- Find if a bacterium is resistant to penicillin (beta-lactamase expression).
Similarly, the assay of the location of protein expression is a powerful tool, and this can be done on an organismal or cellular calibration. Investigation of localization is particularly important for the report of development in multicellular organisms and equally an indicator of protein function in unmarried cells. Ideally, measurement of expression is done by detecting the final gene product (for many genes, this is the protein); however, it is oft easier to detect i of the precursors, typically mRNA and to infer factor-expression levels from these measurements.
mRNA quantification [edit]
Levels of mRNA can be quantitatively measured past northern blotting, which provides size and sequence data almost the mRNA molecules. A sample of RNA is separated on an agarose gel and hybridized to a radioactively labeled RNA probe that is complementary to the target sequence. The radiolabeled RNA is then detected past an autoradiograph. Because the use of radioactive reagents makes the procedure time-consuming and potentially dangerous, alternative labeling and detection methods, such as digoxigenin and biotin chemistries, have been adult. Perceived disadvantages of Northern blotting are that large quantities of RNA are required and that quantification may not be completely accurate, as it involves measuring band strength in an prototype of a gel. On the other hand, the additional mRNA size information from the Northern blot allows the bigotry of alternately spliced transcripts.
Some other approach for measuring mRNA abundance is RT-qPCR. In this technique, reverse transcription is followed past quantitative PCR. Reverse transcription first generates a DNA template from the mRNA; this unmarried-stranded template is chosen cDNA. The cDNA template is then amplified in the quantitative pace, during which the fluorescence emitted past labeled hybridization probes or intercalating dyes changes equally the Dna amplification process progresses. With a carefully synthetic standard curve, qPCR tin produce an accented measurement of the number of copies of original mRNA, typically in units of copies per nanolitre of homogenized tissue or copies per jail cell. qPCR is very sensitive (detection of a unmarried mRNA molecule is theoretically possible), but can be expensive depending on the type of reporter used; fluorescently labeled oligonucleotide probes are more expensive than non-specific intercalating fluorescent dyes.
For expression profiling, or high-throughput assay of many genes within a sample, quantitative PCR may be performed for hundreds of genes simultaneously in the case of low-density arrays. A second approach is the hybridization microarray. A single array or "flake" may contain probes to determine transcript levels for every known cistron in the genome of one or more organisms. Alternatively, "tag based" technologies like Serial analysis of cistron expression (SAGE) and RNA-Seq, which tin can provide a relative measure of the cellular concentration of different mRNAs, can be used. An advantage of tag-based methods is the "open architecture", allowing for the exact measurement of any transcript, with a known or unknown sequence. Adjacent-generation sequencing (NGS) such equally RNA-Seq is another approach, producing vast quantities of sequence data that can be matched to a reference genome. Although NGS is insufficiently time-consuming, expensive, and resource-intensive, information technology can identify single-nucleotide polymorphisms, splice-variants, and novel genes, and can also be used to contour expression in organisms for which trivial or no sequence data is bachelor.
RNA profiles in Wikipedia [edit]
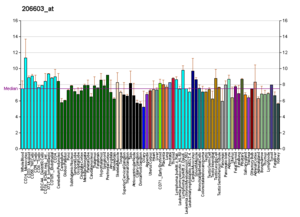
The RNA expression profile of the GLUT4 Transporter (1 of the main glucose transporters constitute in the human body)
Profiles like these are found for well-nigh all proteins listed in Wikipedia. They are generated by organizations such as the Genomics Found of the Novartis Enquiry Foundation and the European Bioinformatics Institute. Additional data can be establish past searching their databases (for an example of the GLUT4 transporter pictured here, see commendation).[75] These profiles indicate the level of Dna expression (and hence RNA produced) of a certain poly peptide in a certain tissue, and are color-coded appropriately in the images located in the Protein Box on the right side of each Wikipedia folio.
Protein quantification [edit]
For genes encoding proteins, the expression level can be direct assessed by a number of methods with some clear analogies to the techniques for mRNA quantification.
One of the well-nigh commonly used methods is to perform a Western blot against the protein of involvement.[76] This gives information on the size of the protein in add-on to its identity. A sample (often cellular lysate) is separated on a polyacrylamide gel, transferred to a membrane and then probed with an antibiotic to the protein of interest. The antibody can either exist conjugated to a fluorophore or to horseradish peroxidase for imaging and/or quantification. The gel-based nature of this analysis makes quantification less authentic, but it has the advantage of beingness able to identify later modifications to the poly peptide, for example proteolysis or ubiquitination, from changes in size.
mRNA-protein correlation [edit]
Quantification of protein and mRNA permits a correlation of the ii levels. The question of how well protein levels correlate with their respective transcript levels is highly debated and depends on multiple factors. Regulation on each step of gene expression can impact the correlation, equally shown for regulation of translation[19] or protein stability.[77] Postal service-translational factors, such equally protein transport in highly polar cells,[78] can influence the measured mRNA-protein correlation as well.
Localisation [edit]
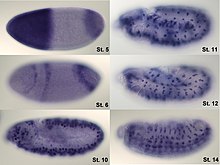
In situ-hybridization of Drosophila embryos at unlike developmental stages for the mRNA responsible for the expression of hunchback. High intensity of blue color marks places with high hunchback mRNA quantity.
Analysis of expression is not limited to quantification; localisation tin can as well be determined. mRNA can exist detected with a suitably labelled complementary mRNA strand and protein tin can be detected via labelled antibodies. The probed sample is then observed past microscopy to identify where the mRNA or protein is.
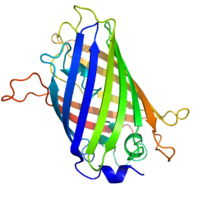
The three-dimensional construction of dark-green fluorescent protein. The residues in the centre of the "barrel" are responsible for production of dark-green light after exposing to higher energetic bluish lite. From PDB: 1EMA.
By replacing the cistron with a new version fused to a green fluorescent protein (or like) mark, expression may be directly quantified in live cells. This is done past imaging using a fluorescence microscope. It is very hard to clone a GFP-fused protein into its native location in the genome without affecting expression levels so this method often cannot be used to mensurate endogenous gene expression. Information technology is, still, widely used to measure the expression of a factor artificially introduced into the cell, for example via an expression vector. Information technology is of import to note that by fusing a target protein to a fluorescent reporter the protein'south behavior, including its cellular localization and expression level, can be significantly changed.
The enzyme-linked immunosorbent assay works past using antibodies immobilised on a microtiter plate to capture proteins of involvement from samples added to the well. Using a detection antibody conjugated to an enzyme or fluorophore the quantity of leap protein can be accurately measured by fluorometric or colourimetric detection. The detection procedure is very similar to that of a Western absorb, only by avoiding the gel steps more accurate quantification tin be achieved.
Expression system [edit]
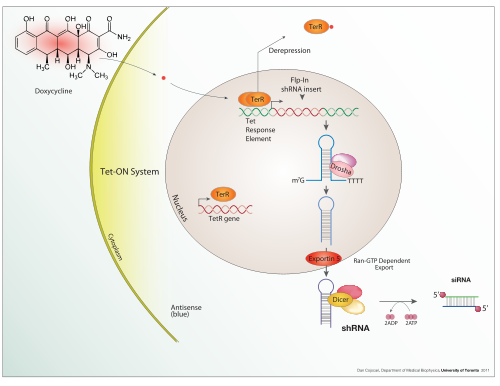
Tet-ON inducible shRNA system
An expression system is a organisation specifically designed for the product of a gene product of choice. This is ordinarily a protein although may as well be RNA, such equally tRNA or a ribozyme. An expression system consists of a cistron, normally encoded by Dna, and the molecular mechanism required to transcribe the Deoxyribonucleic acid into mRNA and interpret the mRNA into protein using the reagents provided. In the broadest sense this includes every living cell but the term is more than normally used to refer to expression every bit a laboratory tool. An expression arrangement is therefore frequently artificial in some manner. Expression systems are, however, a fundamentally natural process. Viruses are an excellent example where they replicate by using the host cell equally an expression system for the viral proteins and genome.
Inducible expression [edit]
Doxycycline is also used in "Tet-on" and "Tet-off" tetracycline controlled transcriptional activation to regulate transgene expression in organisms and cell cultures.
In nature [edit]
In add-on to these biological tools, certain naturally observed configurations of Deoxyribonucleic acid (genes, promoters, enhancers, repressors) and the associated machinery itself are referred to equally an expression system. This term is normally used in the case where a gene or gear up of genes is switched on nether well divers atmospheric condition, for example, the simple repressor switch expression system in Lambda phage and the lac operator organisation in bacteria. Several natural expression systems are directly used or modified and used for artificial expression systems such as the Tet-on and Tet-off expression system.
Gene networks [edit]
Genes accept sometimes been regarded equally nodes in a network, with inputs existence proteins such every bit transcription factors, and outputs being the level of gene expression. The node itself performs a part, and the functioning of these functions take been interpreted every bit performing a kind of information processing inside cells and determines cellular beliefs.
Gene networks tin can besides be constructed without formulating an explicit causal model. This is often the case when assembling networks from large expression data sets.[79] Covariation and correlation of expression is computed beyond a large sample of cases and measurements (often transcriptome or proteome data). The source of variation can be either experimental or natural (observational). There are several means to construct cistron expression networks, but one common arroyo is to compute a matrix of all pair-wise correlations of expression across weather, time points, or individuals and catechumen the matrix (subsequently thresholding at some cut-off value) into a graphical representation in which nodes stand for genes, transcripts, or proteins and edges connecting these nodes represent the forcefulness of association (see [ane]).[80]
Techniques and tools [edit]
The post-obit experimental techniques are used to measure cistron expression and are listed in roughly chronological order, starting with the older, more established technologies. They are divided into 2 groups based on their degree of multiplexity.
- Low-to-mid-plex techniques:
- Reporter gene
- Northern absorb
- Western blot[81]
- Fluorescent in situ hybridization
- Reverse transcription PCR
- Higher-plex techniques:
- SAGE[82]
- Deoxyribonucleic acid microarray[83]
- Tiling array[84]
- RNA-Seq[85]
Gene expression databases [edit]
- Gene expression omnibus (GEO) at NCBI[86]
- Expression Atlas at the EBI
- Mouse Factor Expression Database at the Jackson Laboratory
- CollecTF: a database of experimentally validated transcription cistron-bounden sites in Bacteria.[87]
- COLOMBOS: collection of bacterial expression compendia.[88]
- Many Microbe Microarrays Database: microbial Affymetrix data[89]
See also [edit]
- AlloMap molecular expression testing
- Bookmarking
- Expressed sequence tag
- Expression Atlas
- Expression profiling
- Gene construction
- Genetic engineering
- Genetically modified organism
- List of biological databases
- List of human genes
- Aquiver gene
- Paramutation
- Protein production
- Protein purification
- Ribonomics
- Ridge
- Sequence profiling tool
- Transcriptional bursting
- Transcriptional dissonance
- Transcript of unknown role
References [edit]
- ^ Crick FH (1958). "On protein synthesis". Symposia of the Society for Experimental Biology. 12: 138–63. PMID 13580867.
- ^ Crick F (August 1970). "Fundamental dogma of molecular biological science". Nature. 227 (5258): 561–3. Bibcode:1970Natur.227..561C. doi:10.1038/227561a0. PMID 4913914. S2CID 4164029.
- ^ "Central dogma reversed". Nature. 226 (5252): 1198–ix. June 1970. Bibcode:1970Natur.226.1198.. doi:10.1038/2261198a0. PMID 5422595. S2CID 4184060.
- ^ Temin HM, Mizutani Due south (June 1970). "RNA-dependent Deoxyribonucleic acid polymerase in virions of Rous sarcoma virus". Nature. 226 (5252): 1211–3. doi:10.1038/2261211a0. PMID 4316301. S2CID 4187764.
- ^ Baltimore D (June 1970). "RNA-dependent DNA polymerase in virions of RNA tumour viruses". Nature. 226 (5252): 1209–11. doi:x.1038/2261209a0. PMID 4316300. S2CID 4222378.
- ^ Iyer LM, Koonin EV, Aravind L (January 2003). "Evolutionary connection between the catalytic subunits of DNA-dependent RNA polymerases and eukaryotic RNA-dependent RNA polymerases and the origin of RNA polymerases". BMC Structural Biology. iii: 1. doi:10.1186/1472-6807-3-1. PMC151600. PMID 12553882.
- ^ Brueckner F, Armache KJ, Cheung A, Damsma GE, Kettenberger H, Lehmann E, Sydow J, Cramer P (February 2009). "Structure-part studies of the RNA polymerase Ii elongation complex". Acta Crystallographica D. 65 (Pt 2): 112–xx. doi:10.1107/S0907444908039875. PMC2631633. PMID 19171965.
- ^ Krebs, Jocelyn E. (2017-03-02). Lewin's genes XII. Goldstein, Elliott S.,, Kilpatrick, Stephen T. Burlington, MA. ISBN978-ane-284-10449-3. OCLC 965781334.
- ^ Sirri V, Urcuqui-Inchima Due south, Roussel P, Hernandez-Verdun D (January 2008). "Nucleolus: the fascinating nuclear trunk". Histochemistry and Prison cell Biology. 129 (one): 13–31. doi:x.1007/s00418-007-0359-half-dozen. PMC2137947. PMID 18046571.
- ^ Frank DN, Pace NR (1998). "Ribonuclease P: unity and variety in a tRNA processing ribozyme". Annual Review of Biochemistry. 67: 153–80. doi:10.1146/annurev.biochem.67.one.153. PMID 9759486.
- ^ Ceballos M, Vioque A (2007). "tRNase Z". Poly peptide and Peptide Letters. 14 (2): 137–45. doi:10.2174/092986607779816050. PMID 17305600.
- ^ Weiner AM (October 2004). "tRNA maturation: RNA polymerization without a nucleic acid template". Current Biological science. 14 (20): R883–5. doi:x.1016/j.cub.2004.09.069. PMID 15498478.
- ^ Köhler A, Hurt E (Oct 2007). "Exporting RNA from the nucleus to the cytoplasm". Nature Reviews. Molecular Cell Biological science. 8 (x): 761–73. doi:ten.1038/nrm2255. PMID 17786152. S2CID 10836137.
- ^ Jambhekar A, Derisi JL (May 2007). "Cis-acting determinants of asymmetric, cytoplasmic RNA transport". RNA. 13 (5): 625–42. doi:x.1261/rna.262607. PMC1852811. PMID 17449729.
- ^ Amaral PP, Dinger ME, Mercer TR, Mattick JS (March 2008). "The eukaryotic genome equally an RNA machine". Scientific discipline. 319 (5871): 1787–9. Bibcode:2008Sci...319.1787A. doi:ten.1126/science.1155472. PMID 18369136. S2CID 206511756.
- ^ Hansen TM, Baranov PV, Ivanov IP, Gesteland RF, Atkins JF (May 2003). "Maintenance of the correct open reading frame by the ribosome". EMBO Reports. four (5): 499–504. doi:10.1038/sj.embor.embor825. PMC1319180. PMID 12717454.
- ^ Berk V, Cate JH (June 2007). "Insights into protein biosynthesis from structures of bacterial ribosomes". Current Stance in Structural Biological science. 17 (3): 302–9. doi:10.1016/j.sbi.2007.05.009. PMID 17574829.
- ^ Schwanhäusser B, Busse D, Li North, Dittmar G, Schuchhardt J, Wolf J, Chen W, Selbach M (May 2011). "Global quantification of mammalian cistron expression command" (PDF). Nature. 473 (7347): 337–42. Bibcode:2011Natur.473..337S. doi:10.1038/nature10098. PMID 21593866. S2CID 205224972.
- ^ a b Schwanhäusser B, Busse D, Li N, Dittmar K, Schuchhardt J, Wolf J, Chen W, Selbach M (March 2013). "Blunder: Global quantification of mammalian factor expression control". Nature. 495 (7439): 126–7. Bibcode:2013Natur.495..126S. doi:10.1038/nature11848. PMID 23407496.
- ^ Hegde RS, Kang SW (July 2008). "The concept of translocational regulation". The Journal of Cell Biology. 182 (2): 225–32. doi:10.1083/jcb.200804157. PMC2483521. PMID 18644895.
- ^ Alberts B, Johnson A, Lewis J, Raff G, Roberts Thousand, Walters P (2002). "The Shape and Structure of Proteins". Molecular Biology of the Jail cell; Quaternary Edition. New York and London: Garland Science. ISBN978-0-8153-3218-3.
- ^ Anfinsen CB (July 1972). "The formation and stabilization of poly peptide structure". The Biochemical Journal. 128 (iv): 737–49. doi:10.1042/bj1280737. PMC1173893. PMID 4565129.
- ^ Jeremy Grand. Berg, John 50. Tymoczko, Lubert Stryer; Web content by Neil D. Clarke (2002). "iii. Protein Structure and Function". Biochemistry. San Francisco: W. H. Freeman. ISBN978-0-7167-4684-iii.
{{cite book}}: CS1 maint: multiple names: authors list (link) - ^ Selkoe DJ (December 2003). "Folding proteins in fatal ways". Nature. 426 (6968): 900–4. Bibcode:2003Natur.426..900S. doi:x.1038/nature02264. PMID 14685251. S2CID 6451881.
- ^ Alberts B, Bray D, Hopkin K, Johnson A, Lewis J, Raff One thousand, Roberts G, Walter P (2010). "Protein Structure and Role". Essential Cell Biology (3rd ed.). New York: Garland Science, Taylor and Francis Group, LLC. pp. 120–170. ISBN978-0815341291.
- ^ Hebert DN, Molinari G (October 2007). "In and out of the ER: protein folding, quality control, deposition, and related man diseases". Physiological Reviews. 87 (4): 1377–408. doi:x.1152/physrev.00050.2006. PMID 17928587.
- ^ Russell R (Jan 2008). "RNA misfolding and the action of chaperones". Frontiers in Bioscience. 13 (thirteen): 1–20. doi:x.2741/2557. PMC2610265. PMID 17981525.
- ^ Kober L, Zehe C, Bode J (April 2013). "Optimized indicate peptides for the development of high expressing CHO prison cell lines". Biotechnology and Bioengineering. 110 (4): 1164–73. doi:ten.1002/flake.24776. PMID 23124363. S2CID 449870.
- ^ Moreau P, Brandizzi F, Hanton S, Chatre L, Melser S, Hawes C, Satiat-Jeunemaitre B (2007). "The constitute ER-Golgi interface: a highly structured and dynamic membrane circuitous". Journal of Experimental Botany. 58 (1): 49–64. doi:10.1093/jxb/erl135. PMID 16990376.
- ^ Prudovsky I, Tarantini F, Landriscina M, Neivandt D, Soldi R, Kirov A, Small D, Kathir KM, Rajalingam D, Kumar TK (April 2008). "Secretion without Golgi". Journal of Cellular Biochemistry. 103 (5): 1327–43. doi:10.1002/jcb.21513. PMC2613191. PMID 17786931.
- ^ Zaidi SK, Young DW, Choi JY, Pratap J, Javed A, Montecino M, Stein JL, Lian JB, van Wijnen AJ, Stein GS (Oct 2004). "Intranuclear trafficking: organization and assembly of regulatory machinery for combinatorial biological command". The Periodical of Biological Chemistry. 279 (42): 43363–vi. doi:10.1074/jbc.R400020200. PMID 15277516.
- ^ Mattick JS, Amaral PP, Dinger ME, Mercer TR, Mehler MF (January 2009). "RNA regulation of epigenetic processes". BioEssays. 31 (i): 51–ix. doi:10.1002/bies.080099. PMID 19154003.
- ^ Martinez NJ, Walhout AJ (Apr 2009). "The coaction between transcription factors and microRNAs in genome-scale regulatory networks". BioEssays. 31 (4): 435–45. doi:10.1002/bies.200800212. PMC3118512. PMID 19274664.
- ^ Tomilin NV (April 2008). "Regulation of mammalian gene expression past retroelements and not-coding tandem repeats". BioEssays. 30 (iv): 338–48. doi:10.1002/bies.20741. PMID 18348251.
- ^ Veitia RA (November 2008). "1 k and i ways of making functionally similar transcriptional enhancers". BioEssays. xxx (11–12): 1052–vii. doi:10.1002/bies.20849. PMID 18937349.
- ^ Nguyen T, Nioi P, Pickett CB (May 2009). "The Nrf2-antioxidant response element signaling pathway and its activation by oxidative stress". The Periodical of Biological Chemistry. 284 (20): 13291–5. doi:10.1074/jbc.R900010200. PMC2679427. PMID 19182219.
- ^ Paul S (November 2008). "Dysfunction of the ubiquitin-proteasome system in multiple illness conditions: therapeutic approaches". BioEssays. 30 (11–12): 1172–84. doi:10.1002/bies.20852. PMID 18937370. S2CID 29422790.
- ^ Los Thou, Maddika Southward, Erb B, Schulze-Osthoff K (May 2009). "Switching Akt: from survival signaling to deadly response". BioEssays. 31 (five): 492–five. doi:10.1002/bies.200900005. PMC2954189. PMID 19319914.
- ^ Verheul TC, van Hijfte L, Perenthaler E, Barakat TS (2020). "The Why of YY1: Mechanisms of Transcriptional Regulation by Yin Yang 1". Front Cell Dev Biol. 8: 592164. doi:10.3389/fcell.2020.592164. PMC7554316. PMID 33102493.
- ^ Spitz F, Furlong EE (September 2012). "Transcription factors: from enhancer binding to developmental command". Nat Rev Genet. 13 (ix): 613–26. doi:x.1038/nrg3207. PMID 22868264. S2CID 205485256.
- ^ a b Schoenfelder S, Fraser P (August 2019). "Long-range enhancer-promoter contacts in cistron expression control". Nat Rev Genet. 20 (eight): 437–455. doi:10.1038/s41576-019-0128-0. PMID 31086298. S2CID 152283312.
- ^ Weintraub Equally, Li CH, Zamudio AV, Sigova AA, Hannett NM, Day DS, Abraham BJ, Cohen MA, Nabet B, Buckley DL, Guo YE, Hnisz D, Jaenisch R, Bradner JE, Grey NS, Young RA (Dec 2017). "YY1 Is a Structural Regulator of Enhancer-Promoter Loops". Cell. 171 (7): 1573–1588.e28. doi:10.1016/j.jail cell.2017.11.008. PMC5785279. PMID 29224777.
- ^ Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, Chen X, Taipale J, Hughes TR, Weirauch MT (Feb 2018). "The Human Transcription Factors". Jail cell. 172 (4): 650–665. doi:10.1016/j.cell.2018.01.029. PMID 29425488.
- ^ Grossman SR, Engreitz J, Ray JP, Nguyen Thursday, Hacohen N, Lander ES (July 2018). "Positional specificity of different transcription factor classes within enhancers". Proc Natl Acad Sci U S A. 115 (30): E7222–E7230. doi:ten.1073/pnas.1804663115. PMC6065035. PMID 29987030.
- ^ Allen BL, Taatjes DJ (March 2015). "The Mediator complex: a fundamental integrator of transcription". Nat Rev Mol Cell Biol. 16 (iii): 155–66. doi:10.1038/nrm3951. PMC4963239. PMID 25693131.
- ^ Mikhaylichenko O, Bondarenko V, Harnett D, Schor IE, Males M, Viales RR, Furlong EE (Jan 2018). "The degree of enhancer or promoter action is reflected by the levels and directionality of eRNA transcription". Genes Dev. 32 (one): 42–57. doi:10.1101/gad.308619.117. PMC5828394. PMID 29378788.
- ^ Li QJ, Yang SH, Maeda Y, Sladek FM, Sharrocks Ad, Martins-Greenish Grand (January 2003). "MAP kinase phosphorylation-dependent activation of Elk-1 leads to activation of the co-activator p300". EMBO J. 22 (2): 281–91. doi:10.1093/emboj/cdg028. PMC140103. PMID 12514134.
- ^ Carullo NV, Phillips I RA, Simon RC, Soto SA, Hinds JE, Salisbury AJ, Revanna JS, Bunner KD, Ianov L, Sultan FA, Savell KE, Gersbach CA, Day JJ (September 2020). "Enhancer RNAs predict enhancer-factor regulatory links and are critical for enhancer office in neuronal systems". Nucleic Acids Res. 48 (17): 9550–9570. doi:10.1093/nar/gkaa671. PMC7515708. PMID 32810208.
- ^ Lövkvist C, Dodd IB, Sneppen G, Haerter JO (June 2016). "Dna methylation in human epigenomes depends on local topology of CpG sites". Nucleic Acids Research. 44 (11): 5123–32. doi:x.1093/nar/gkw124. PMC4914085. PMID 26932361.
- ^ Jabbari K, Bernardi G (May 2004). "Cytosine methylation and CpG, TpG (CpA) and TpA frequencies". Cistron. 333: 143–9. doi:10.1016/j.gene.2004.02.043. PMID 15177689.
- ^ Weber Thousand, Hellmann I, Stadler MB, Ramos Fifty, Pääbo S, Rebhan M, Schübeler D (April 2007). "Distribution, silencing potential and evolutionary bear on of promoter Deoxyribonucleic acid methylation in the human genome". Nat. Genet. 39 (four): 457–66. doi:10.1038/ng1990. PMID 17334365. S2CID 22446734.
- ^ Yang X, Han H, De Carvalho DD, Lay FD, Jones PA, Liang G (October 2014). "Gene body methylation tin alter gene expression and is a therapeutic target in cancer". Cancer Cell. 26 (four): 577–ninety. doi:10.1016/j.ccr.2014.07.028. PMC4224113. PMID 25263941.
- ^ Maeder ML, Angstman JF, Richardson ME, Linder SJ, Cascio VM, Tsai SQ, Ho QH, Sander JD, Reyon D, Bernstein BE, Costello JF, Wilkinson MF, Joung JK (December 2013). "Targeted DNA demethylation and activation of endogenous genes using programmable TALE-TET1 fusion proteins". Nat. Biotechnol. 31 (12): 1137–42. doi:10.1038/nbt.2726. PMC3858462. PMID 24108092.
- ^ Kim JJ, Jung MW (2006). "Neural circuits and mechanisms involved in Pavlovian fear conditioning: a critical review". Neuroscience and Biobehavioral Reviews. 30 (two): 188–202. doi:10.1016/j.neubiorev.2005.06.005. PMC4342048. PMID 16120461.
- ^ a b Knuckles CG, Kennedy AJ, Gavin CF, Solar day JJ, Sweatt JD (July 2017). "Experience-dependent epigenomic reorganization in the hippocampus". Learning & Memory. 24 (7): 278–288. doi:10.1101/lm.045112.117. PMC5473107. PMID 28620075.
- ^ a b Keifer J (Feb 2017). "Primetime for Learning Genes". Genes (Basel). 8 (2): 69. doi:10.3390/genes8020069. PMC5333058. PMID 28208656.
- ^ Saxonov S, Berg P, Brutlag DL (Jan 2006). "A genome-wide assay of CpG dinucleotides in the human being genome distinguishes ii distinct classes of promoters". Proceedings of the National University of Sciences of the Usa of America. 103 (5): 1412–vii. Bibcode:2006PNAS..103.1412S. doi:10.1073/pnas.0510310103. PMC1345710. PMID 16432200.
- ^ Bird A (January 2002). "DNA methylation patterns and epigenetic retention". Genes & Development. 16 (1): 6–21. doi:ten.1101/gad.947102. PMID 11782440.
- ^ Vogelstein B, Papadopoulos N, Velculescu VE, Zhou Southward, Diaz LA, Kinzler KW (March 2013). "Cancer genome landscapes". Science. 339 (6127): 1546–58. Bibcode:2013Sci...339.1546V. doi:10.1126/science.1235122. PMC3749880. PMID 23539594.
- ^ Tessitore A, Cicciarelli G, Del Vecchio F, Gaggiano A, Verzella D, Fischietti Grand, Vecchiotti D, Capece D, Zazzeroni F, Alesse East (2014). "MicroRNAs in the DNA Damage/Repair Network and Cancer". International Journal of Genomics. 2014: 1–10. doi:x.1155/2014/820248. PMC3926391. PMID 24616890.
- ^ miRBase.org
- ^ a b Friedman RC, Farh KK, Burge CB, Bartel DP (Jan 2009). "Most mammalian mRNAs are conserved targets of microRNAs". Genome Enquiry. xix (1): 92–105. doi:10.1101/gr.082701.108. PMC2612969. PMID 18955434.
- ^ Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, Johnson JM (February 2005). "Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs". Nature. 433 (7027): 769–73. Bibcode:2005Natur.433..769L. doi:10.1038/nature03315. PMID 15685193. S2CID 4430576.
- ^ Selbach Yard, Schwanhäusser B, Thierfelder N, Fang Z, Khanin R, Rajewsky Northward (September 2008). "Widespread changes in protein synthesis induced by microRNAs". Nature. 455 (7209): 58–63. Bibcode:2008Natur.455...58S. doi:10.1038/nature07228. PMID 18668040. S2CID 4429008.
- ^ Baek D, Villén J, Shin C, Camargo FD, Gygi SP, Bartel DP (September 2008). "The touch of microRNAs on protein output". Nature. 455 (7209): 64–71. Bibcode:2008Natur.455...64B. doi:x.1038/nature07242. PMC2745094. PMID 18668037.
- ^ Palmero EI, de Campos SG, Campos Grand, de Souza NC, Guerreiro ID, Carvalho AL, Marques MM (July 2011). "Mechanisms and part of microRNA deregulation in cancer onset and progression". Genetics and Molecular Biology. 34 (3): 363–70. doi:10.1590/S1415-47572011000300001. PMC3168173. PMID 21931505.
- ^ Bernstein C, Bernstein H (May 2015). "Epigenetic reduction of Dna repair in progression to gastrointestinal cancer". World Journal of Gastrointestinal Oncology. 7 (5): xxx–46. doi:10.4251/wjgo.v7.i5.thirty. PMC4434036. PMID 25987950.
- ^ Mellios N, Sur M (2012). "The Emerging Role of microRNAs in Schizophrenia and Autism Spectrum Disorders". Frontiers in Psychiatry. 3: 39. doi:10.3389/fpsyt.2012.00039. PMC3336189. PMID 22539927.
- ^ Geaghan M, Cairns MJ (Baronial 2015). "MicroRNA and Posttranscriptional Dysregulation in Psychiatry". Biological Psychiatry. 78 (four): 231–9. doi:10.1016/j.biopsych.2014.12.009. PMID 25636176.
- ^ a b c Walsh CT, Garneau-Tsodikova Southward, Gatto GJ (Dec 2005). "Protein posttranslational modifications: the chemistry of proteome diversifications". Angewandte Chemie. 44 (45): 7342–72. doi:10.1002/anie.200501023. PMID 16267872. S2CID 32157563.
- ^ Khoury GA, Baliban RC, Floudas CA (September 2011). "Proteome-wide post-translational modification statistics: frequency assay and curation of the swiss-prot database". Scientific Reports. 1 (xc): 90. Bibcode:2011NatSR...1E..90K. doi:10.1038/srep00090. PMC3201773. PMID 22034591.
- ^ Mann M, Jensen ON (March 2003). "Proteomic analysis of mail-translational modifications". Nature Biotechnology. 21 (iii): 255–61. doi:10.1038/nbt0303-255. PMID 12610572. S2CID 205266061.
- ^ Seo J, Lee KJ (January 2004). "Postal service-translational modifications and their biological functions: proteomic analysis and systematic approaches". Periodical of Biochemistry and Molecular Biology. 37 (1): 35–44. doi:10.5483/bmbrep.2004.37.one.035. PMID 14761301.
- ^ Rogers LD, Overall CM (December 2013). "Proteolytic post-translational modification of proteins: proteomic tools and methodology". Molecular & Cellular Proteomics. 12 (12): 3532–42. doi:10.1074/mcp.M113.031310. PMC3861706. PMID 23887885.
- ^ "GLUT4 RNA Expression Profile".
- ^ Moritz, CP (10 February 2020). "40 years Western blotting: A scientific birthday toast". Journal of Proteomics. 212: 103575. doi:10.1016/j.jprot.2019.103575. PMID 31706026.
- ^ Burkhart JM, Vaudel M, Gambaryan S, Radau S, Walter U, Martens 50, Geiger J, Sickmann A, Zahedi RP (October 2011). "The start comprehensive and quantitative analysis of human platelet protein composition allows the comparative analysis of structural and functional pathways". Blood. 120 (15): e73–82. doi:10.1182/blood-2012-04-416594. PMID 22869793.
- ^ Moritz CP, Mühlhaus T, Tenzer Due south, Schulenborg T, Friauf E (June 2019). "Poor transcript-protein correlation in the encephalon: negatively correlating cistron products reveal neuronal polarity as a potential crusade" (PDF). Journal of Neurochemistry. 149 (5): 582–604. doi:x.1111/jnc.14664. PMID 30664243. S2CID 58667771.
- ^ Banf G, Rhee SY (January 2017). "Computational inference of factor regulatory networks: Approaches, limitations and opportunities". Biochimica et Biophysica Acta (BBA) - Factor Regulatory Mechanisms. 1860 (1): 41–52. doi:10.1016/j.bbagrm.2016.09.003. PMID 27641093.
- ^ Chesler EJ, Lu Fifty, Wang J, Williams RW, Manly KF (May 2004). "WebQTL: rapid exploratory assay of factor expression and genetic networks for brain and behavior". Nature Neuroscience. 7 (5): 485–6. doi:10.1038/nn0504-485. PMID 15114364. S2CID 20241963.
- ^ Song Y, Wang W, Qu X, Sun S (Feb 2009). "Effects of hypoxia inducible factor-1alpha (HIF-1alpha) on the growth & adhesion in tongue squamous cell carcinoma cells". The Indian Journal of Medical Inquiry. 129 (2): 154–63. PMID 19293442.
- ^ Hanriot 50, Keime C, Gay Due north, Faure C, Dossat C, Wincker P, Scoté-Blachon C, Peyron C, Gandrillon O (September 2008). "A combination of LongSAGE with Solexa sequencing is well suited to explore the depth and the complexity of transcriptome". BMC Genomics. 9: 418. doi:x.1186/1471-2164-9-418. PMC2562395. PMID 18796152.
- ^ Wheelan SJ, Martínez Murillo F, Boeke JD (July 2008). "The incredible shrinking earth of DNA microarrays". Molecular BioSystems. four (vii): 726–32. doi:10.1039/b706237k. PMC2535915. PMID 18563246.
- ^ Miyakoshi G, Nishida H, Shintani M, Yamane H, Nojiri H (Jan 2009). "Loftier-resolution mapping of plasmid transcriptomes in different host leaner". BMC Genomics. ten: 12. doi:10.1186/1471-2164-10-12. PMC2642839. PMID 19134166.
- ^ Denoeud F, Aury JM, Da Silva C, Noel B, Rogier O, Delledonne M, Morgante M, Valle G, Wincker P, Scarpelli C, Jaillon O, Artiguenave F (2008). "Annotating genomes with massive-scale RNA sequencing". Genome Biological science. 9 (12): R175. doi:10.1186/gb-2008-9-12-r175. PMC2646279. PMID 19087247.
- ^ Clough East, Barrett T (2016). "The Factor Expression Omnibus Database". Statistical Genomics. Methods in Molecular Biology. Vol. 1418. pp. 93–110. doi:10.1007/978-ane-4939-3578-9_5. ISBN978-i-4939-3576-five. PMC4944384. PMID 27008011.
- ^ Kiliç South, White ER, Sagitova DM, Cornish JP, Erill I (January 2014). "CollecTF: a database of experimentally validated transcription gene-bounden sites in Bacteria". Nucleic Acids Enquiry. 42 (Database effect): D156–60. doi:10.1093/nar/gkt1123. PMC3965012. PMID 24234444.
- ^ Moretto Thousand, Sonego P, Dierckxsens N, Brilli Grand, Bianco L, Ledezma-Tejeida D, et al. (January 2016). "COLOMBOS v3.0: leveraging gene expression compendia for cross-species analyses". Nucleic Acids Research. 44 (D1): D620–3. doi:x.1093/nar/gkv1251. PMC4702885. PMID 26586805.
- ^ Faith JJ, Driscoll ME, Fusaro VA, Cosgrove EJ, Hayete B, Juhn FS, et al. (Jan 2008). "Many Microbe Microarrays Database: uniformly normalized Affymetrix compendia with structured experimental metadata". Nucleic Acids Inquiry. 36 (Database issue): D866–seventy. doi:10.1093/nar/gkm815. PMC2238822. PMID 17932051.
External links [edit]
- Plant Transcription Factor Database and Constitute Transcriptional Regulation Data and Analysis Platform
Source: https://en.wikipedia.org/wiki/Gene_expression
0 Response to "an active gene leads to the production of what? (one word)"
Publicar un comentario